Classification of human epidermal growth factor receptor 2 expression in cancerous breast tissue through artificial intelligence
Abstract
Introduction. Histological and molecular analysis of breast tissue is essential for the diagnosis, prognosis, and treatment of breast cancer. Key biomarkers include progesterone and estrogen receptors, as well as the human epidermal growth factor receptor 2 (HER2). HER2 overexpression indicates an aggressive subtype of breast cancer but enables targeted therapies that improve survival rates. However, its evaluation faces challenges, ranging from sample quality to interpretation variability. The College of American Pathologists classifies HER2 overexpression into four categories, but variations around the 10% expression threshold can lead to misinterpretations.
Objective. To present an automated technique for classifying HER2-overexpressing cells in histological slides.
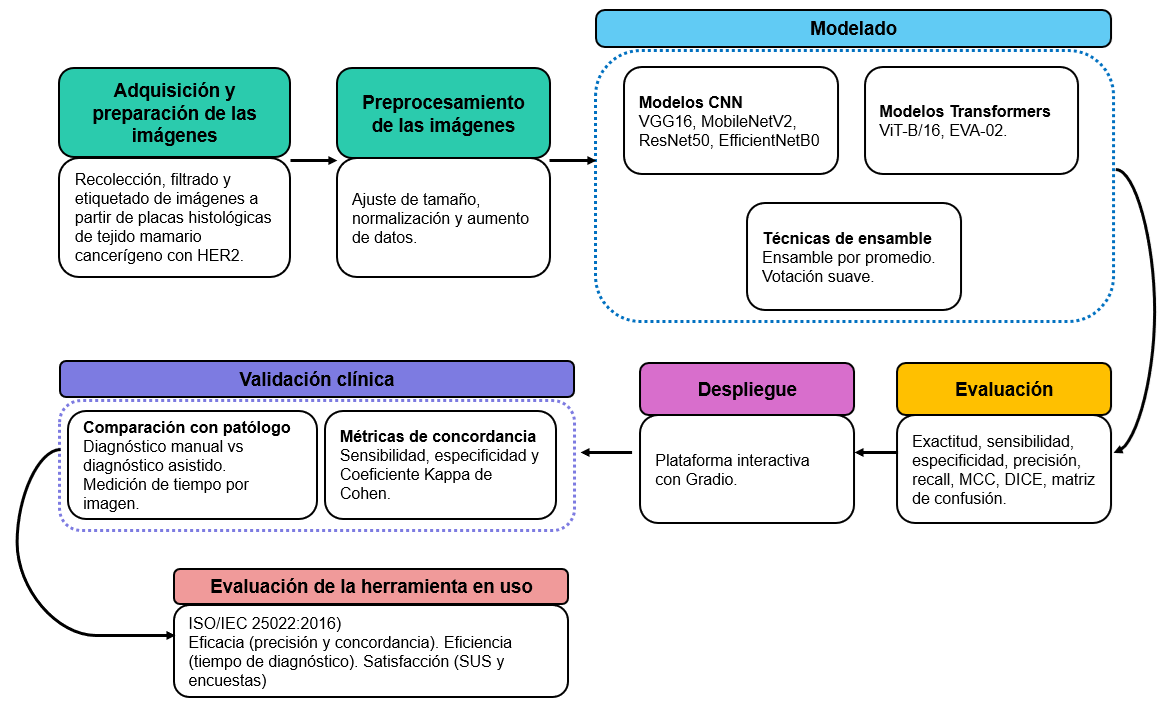
Materials and methods. The Cross-Industry Standard Process for Data Mining (CRISP-DM) methodology was applied using samples of 89 patients from the Unidad de Diagnóstico en Patología, covering all four HER2 expression levels. Deep learning techniques were employed, leveraging neural networks and vision transformer models through transfer learning. Additionally, a usability evaluation was conducted on the final version of the software.
Results. The ViT-B/16 model achieved a classification accuracy of 90,65%, while the tool was evaluated with an acceptable level of satisfaction in its clinical application.
Conclusion. Artificial intelligence demonstrated high accuracy and consistency in HER2 classification, reducing diagnostic variability and improving objectivity. However, further optimization of processing efficiency is required for broader applicability.
Downloads
References
American Cancer Society. What is cancer | Cancer basics. Fecha de consulta: 22 de diciembre de 2024. Disponible en: https://www.cancer.org/cancer/understanding-cancer/what-is-cancer.html
Bórquez S, Pezoa R, Salinas L, Torres CE. Uncertainty estimation in the classification of histopathological images with HER2 overexpression using Monte Carlo Dropout. Biomed Signal Process Control. 2023;85:104864. https://doi.org/10.1016/j.bspc.2023.104864
Swain SM, Shastry M, Hamilton E. Targeting HER2-positive breast cancer: Advances and future directions. Nat Rev Drug Discov. 2023;22:101‑26. https://doi.org/10.1038/s41573-022-00579-0
American Cancer Society. Breast cancer HER2 status | What is HER2 status? Fecha de consulta: 22 de diciembre de 2024. Disponible en: https://www.cancer.org/cancer/types/breast-cancer/understanding-a-breast-cancer-diagnosis/breast-cancer-her2-status.html
American Society of Clinical Oncology. Breast cancer. Fecha de consulta: 22 de diciembre de 2024. Disponible en: https://society.asco.org/practice-patients/guidelines/breast-cancer
International Agency for Research on Cancer. Cancer today. Fecha de consulta: 22 de diciembre de 2024. Disponible en: https://gco.iarc.who.int/today/
Departamento Administrativo Nacional de Estadística. Defunciones no fetales 2020. Fecha de consulta: 22 de diciembre de 2024. Disponible en: https://www.dane.gov.co/index.php/estadisticas-por-tema/salud/nacimientos-y-defunciones/defunciones-no-fetales/defuncionesno-fetales-2020
Zheng Y, Liang H, Zhao S. LMBNet: Lightweight multiple branch network for recognition of HER2 expression levels. Proc Comput Sci. 2023;222:197‑206. https://doi.org/10.1016/j.procs.2023.08.157
Córdova C, Muñoz R, Olivares R, Minonzio JG, Lozano C, González P, et al. HER2 classification in breast cancer cells: A new explainable machine learning application for immunohistochemistry. Oncol Lett. 2023;25:44. https://doi.org/10.3892/ol.2022.13630
Wang X, Shao C, Liu W, Liang H, Li N. HER2‑ResNet: A HER2 classification method based on deep residual network. Technol Health Care. 2022;30(Supl.1):215‑24. https://doi.org/10.3233/THC-228020
Mirimoghaddam MM, Majidpour J, Pashaei F, Arabalibeik H, Samizadeh E, Roshan NM, et al. HER2GAN: Overcome the scarcity of HER2 breast cancer dataset based on transfer learning and GAN Model. Clin Breast Cancer. 2024;24:53‑64. https://doi.org/10.1016/j.clbc.2023.09.014
Selcuk SY, Yang X, Bai B, Zhang Y, Li Y, Aydin M, et al. Automated HER2 scoring in breast cancer images using deep learning and pyramid sampling. BME Front. 2024;5:00‑48. https://doi.org/10.34133/bmef.0048
Chauhan R, Ghanshala KK, Joshi RC. Convolutional Neural Network (CNN) for image detection and recognition. In: First International Conference on Secure Cyber Computing and Communication (ICSCCC); 2018. Jalandhar, India: IEEE; 2018. p. 278‑82. https://doi.org/10.1109/ICSCCC.2018.8703316
Vaswani A, Shazeer N, Parmar N, Uszkoreit J, Jones L, Gómez AN, et al. Attention is all you need. Preprint. arXiv; 2023. arXiv:1706.03762. https://doi.org/10.48550/arXiv.1706.03762
Dosovitskiy A, Beyer L, Kolesnikov A, Weissenborn D, Zhai X, Unterthiner T, et al. An image is worth 16 × 16 words: Transformers for image recognition at scale. Preprint. arXiv; 2021. arXiv:2010.11929. https://doi.org/10.48550/arXiv.2010.11929
Liu Z, Lin Y, Cao Y, Hu H, Wei Y, Zhang Z, et al. Swin transformer: Hierarchical vision transformer using shifted windows. Preprint. arXiv; 2021. arXiv:2103.14030. https://doi.org/10.48550/arXiv.2103.14030
Khan S, Naseer M, Hayat M, Zamir SW, Khan FS, Shah M. Transformers in vision: A survey. ACM Comput Surv. 2022;54(10s):1‑41. https://doi.org/10.1145/3505244
Chollet F. Deep learning with Python. Second edition. New York, NY: Simon and Schuster; 2021. p. 1‑502.
Theckedath D, Sedamkar RR. Detecting affect states using VGG16, ResNet50 and SE‑ResNet50 networks. SN Comput Sci. 2020;1:79. https://doi.org/10.1007/s42979-020-0114-9
Özaltın Ö, Yeniay Ö. Detection of monkeypox disease from skin lesion images using mobilenetv2 architecture. Communications Faculty of Sciences University of Ankara Series A1 Mathematics and Statistics. 2023;72:482‑99. https://doi.org/10.31801/cfsuasmas.1202806
Tan M, Le QV. EfficientNet: Rethinking model scaling for convolutional neural networks. Preprint. arXiv; 2020. arXiv:1905.11946. https://doi.org/10.48550/arXiv.1905.11946
Fang Y, Sun Q, Wang X, Huang T, Wang X, Cao Y. EVA-02: A visual representation for Neon Genesis. Image Vis Comput. 2024;149:105‑71. https://doi.org/10.1016/j.imavis.2024.105171
Stanford Vision Lab; Princeton University. ImageNet. Fecha de consulta: 13 de febrero de 2025. Disponible en: https://www.image-net.org/
Shaga Devan K, Kestler HA, Read C, Walther P. Weighted average ensemble-based semantic segmentation in biological electron microscopy images. Histochem Cell Biol. 2022;158:447‑62. https://doi.org/10.1007/s00418-022-02148-3
Zhou ZH. Ensemble methods: Foundations and algorithms. First edition. Boca Ratón, FL: Chapman & Hall/CRC; 2012. p. 1‑236.
Gradio. Gradio App. Fecha de consulta: 12 de febrero de 2025. Disponible en: https://www.gradio.app/
International Organization for Standardization (ISO). ISO/IEC 25022:2016 — Systems and software engineering — Systems and software quality requirements and evaluation (SQuaRE) - Measurement of quality in use. Fecha de consulta: 12 de febrero de 2025. Disponible en: https://www.iso.org/standard/35746.html
Brooke J. SUS - A quick and dirty usability scale. Fecha de consulta: 12 de febrero de 2025. Disponible en: https://digital.ahrq.gov/sites/default/files/docs/survey/systemusabilityscale%2528sus%2529_comp%255B1%255D.pdf
Some similar items:
- Edwin Abraham Medina, Middle ear adenoma , Biomedica: Vol. 29 No. 3 (2009)
- Aura Caterine Rengifo, Orlando Torres-Fernández, Decreased number neurons expressing GABA in the cerebral cortex of rabies-infected mice , Biomedica: Vol. 27 No. 4 (2007)
- Oscar F. Herrán, María F. Ardila, Martha P. Rojas, Gustavo A. Hernández, Design of dietary questionnaires to study the relationships between diet and cancer prevalence in Colombia , Biomedica: Vol. 30 No. 1 (2010)
- Nina Paola Lamprea, Lina María Ortega, Gerardo Santamaría, Ladys Sarmiento, Orlando Torres-Fernández, Production and evaluation of an antiserum for immunohistochemical detection of rabies virus in aldehyde fixed brain tissues , Biomedica: Vol. 30 No. 1 (2010)
- Juan Carlos Cataño, Eaton-Lambert myasthenic syndrome , Biomedica: Vol. 30 No. 3 (2010)
- Juan Carlos Herrera, Luis Fernando Isaza, José Luis Ramírez, Gonzalo Vásquez, Carlos Mario Muñetón, Detection of chromosome 17 aneuplody and TP53 gene deletion in a broad variety of solid tumors by dual-color fluorescence in situ hybridization (FISH) , Biomedica: Vol. 30 No. 3 (2010)
- Yaliana Tafurt-Cardona, Leidy D. Jaramillo-Ruiz, Wilson Muñoz-Ordóñez, Sulma L. Muñoz-Benítez, Carlos H. Sierra-Torres, High frequency of chromosome aberrations observed in lymphocytes in postmenopausal obese women , Biomedica: Vol. 32 No. 3 (2012)
- Ricardo Cendales, Constanza Pardo, Claudia Uribe, Guillermo López, María Clara Yépez, Luis Eduardo Bravo, Data quality at population-based cancer registries in Colombia , Biomedica: Vol. 32 No. 4 (2012)
- Sonia Isabel Cuervo, Ricardo Sánchez, Julio César Gómez-Rincón, Cielo Almenares, Juan Pablo Osorio, María José Vargas, Behavior of carbapenemase-producing Klebsiella pneumoniae cases in cancer patients at a third level hospital in Bogotá, D.C. , Biomedica: Vol. 34 (2014): Abril, Suplemento 1, Resistencia bacteriana
- Jeison Monroy-Gómez, Orlando Torres-Fernández, Calbindin and parvalbumin distribution in spinal cord of normal and rabies-infected mice , Biomedica: Vol. 33 No. 4 (2013)
Copyright (c) 2025 Biomedica

This work is licensed under a Creative Commons Attribution 4.0 International License.
| Article metrics | |
|---|---|
| Abstract views | |
| Galley vies | |
| PDF Views | |
| HTML views | |
| Other views | |