Use of artificial intelligence in the diagnosis of alterations in cervical cytology: A university population-based observational study
Abstract
Introduction. Conventional cervical cytology (Pap smear) remains a primary method for cervical cancer screening in Colombia, despite limitations in diagnostic yield and heavy workload. The potential of artificial intelligence to address these challenges is yet to be evaluated in our population.
Objective. To evaluate and compare the discriminative ability of four artificial intelligence-based models for the detection of abnormalities in Pap smears.
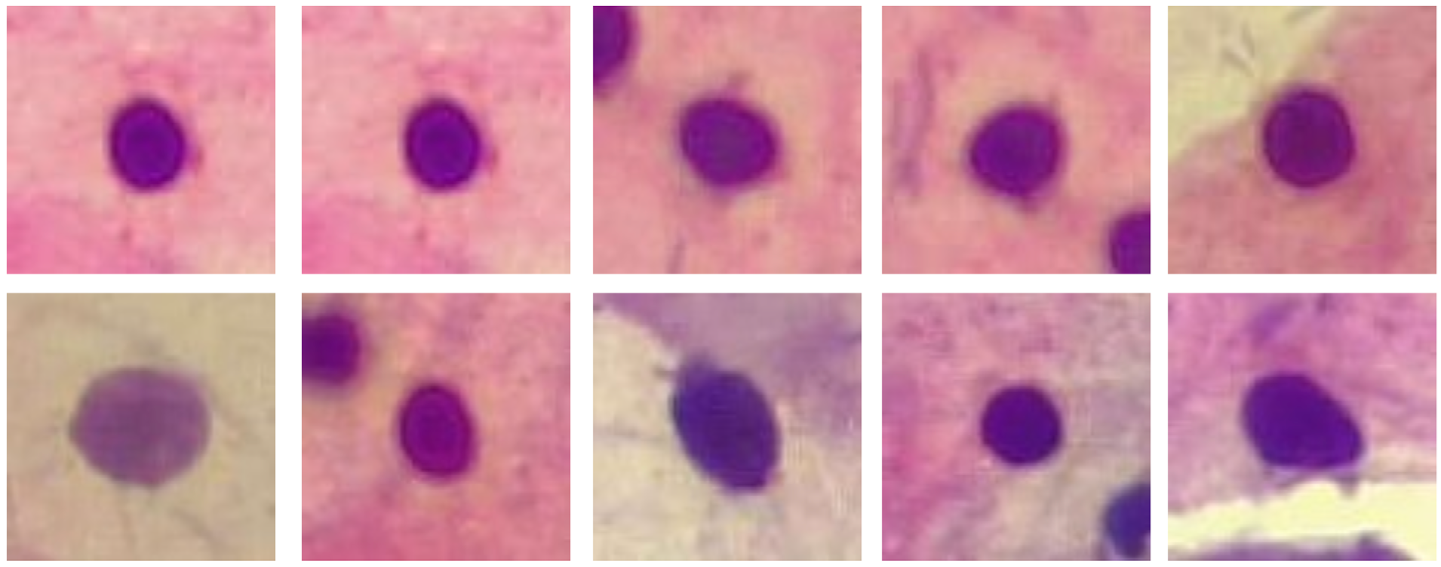
Materials and methods. A total of 650 images of Pap smear cells were obtained from a university cohort in northeastern Colombia. These images were subjected to diagnostic evaluation by an expert pathologist. Four artificial intelligence models (DenseNet, InceptionV3, MobileNet, and VGG19) were trained using data from a publicly available Pap smear database with digital image analysis and deep learning. The discriminative ability of the models was determined by calculating their sensitivity, specificity, and area under the curve.
Results. MobileNet showed the highest discriminative ability (AUC = 0.97), with a specificity of 0.99 and sensitivity of 0.78 for the detection of altered cells in Pap smears. On the other hand, InceptionV3 had the best performance capabilities for screening, with a sensitivity of 0.93, specificity of 0.82, and AUC of 0.947.
Conclusions. The results of this study illustrate the advantages and disadvantages of different artificial intelligence models and how their application could help improve the diagnostic performance of manual reading in cervical cancer screening or even serve as a primary screening method to rule out negative cases, by achieving a diagnostic performance comparable to that of manual reading.
Downloads
References
World Health Organization. Global health estimates 2020: Deaths by cause, age, sex, by country and by region, 2000-2019. Geneva: World Health Organization; 2020.
Organización Panamericana de la Salud. Cáncer cervicouterino - OPS/OMS. Fecha de consulta: 13 de junio de 2022. Disponible en: https://www.paho.org/es/temas/cancercervicouterino
Sanyal P, Ganguli P, Barui S. Performance characteristics of an artificial intelligence based on convolutional neural network for screening conventional Papanicolaou-stained cervical smears. Med J Armed Forces India. 2020;76:418-24. https://doi.org/10.1016/j.mjafi.2019.08.001
Xin M, Wang Y. Research on image classification model based on deep convolution neural network. J Image Video Process. 2019;2019. https://doi.org/10.1186/s13640-019-0417-8
Kumar L, Renuka D, Rose S, Shunmugapriya M, Wartana I. Deep learning based assistive technology on audio visual speech recognition for hearing impaired. Int J Cogn Comput Eng. 2022;3:24-30.
Popel M, Tomkova M, Tomek J, Kaiser Ł, Uszkoreit J, Bojar O, et al. Transforming machine translation: A deep learning system reaches news translation quality comparable to human professionals. Nat Commun. 2020;11:4381. https://doi.org/10.1038/s41467-020-18073-9
Pranav KB, Manikandan J. Design and evaluation of a real-time face recognition system using convolutional neural networks. Procedia Comput Sci. 2020;171:1651-9. https://doi.org/10.1016/j.procs.2020.04.177
Cui M, Zhang DY. Artificial intelligence and computational pathology. Lab Invest. 2021;101:412–22. https://doi.org/10.1038/s41374-020-00514-0
Campanella G, Hanna MG, Geneslaw L, Miraflor A, Werneck KSV, Busam K, et al. Clinicalgrade computational pathology using weakly supervised deep learning on whole slide images. Nat Med. 2019;25:1301-9. https://doi.org/10.1038/s41591-019-0508-1
Korbar B, Olofson AM, Miraflor AP, Nicka CM, Suriawinata MA, Torresani L, et al. Deep learning for classification of colorectal polyps on whole-slide images. J Pathol Inform. 2017;8:30. https://doi.org/10.4103/jpi.jpi_34_17
Bychkov D, Linder N, Turkki R, Nordling S, Kovanen PE, Verrill C, et al. Deep learning based tissue analysis predicts outcome in colorectal cancer. Sci Rep. 2018;8:3395. https://doi.org/10.1038/s41598-018-21758-3
Ehteshami Bejnordi B, Veta M, Johannes van Diest P, Van Ginekken B, Karssemeijer N, Litjens G, et al. Diagnostic assessment of deep learning algorithms for detection of lymph node metastases in women with breast cancer. JAMA. 2017;318:2199-210. https://doi.org/10.1001/jama.2017.14585
Bao H, Sun X, Zhang Y, Pang B, Li H, Zhou L, et al. The artificial intelligence-assisted cytology diagnostic system in large-scale cervical cancer screening: A population-based cohort study of 0.7 million women. Cancer Med. 2020;9:6896-906. https://doi.org/10.1002/cam4.3296
Diniz ND, Rezende MT, Bianchi AG, Carneiro CM, Luz JS, Moreira GP, et al. A deep learning ensemble method to assist cytopathologists in Pap test image classification. J Imaging. 2021;7:111. https://doi.org/10.3390/jimaging7070111
Silva R, Isidoro DWA, Silva AC, Lima SM, Paiva AC. Searching for cell signatures in multidimensional feature spaces. Int J Biomed Eng Technol. 2021;36:236-56.
Rodríguez M, Córdova C, Benjumeda I, San Martín S. Automated cervical cancer screening using single-cell segmentation and deep learning: Enhanced performance with liquid-based cytology. Computation. 2024;12:232. https://doi.org/10.3390/computation12120232
William W, Ware J, Habinka A, Obungoloch J. A review of image analysis and machine learning techniques for automated cervical cancer screening from Pap-smear images. Comput Methods Programs Biomed. 2018;164:15-22. https://doi.org/10.1016/j.cmpb.2018.05.034
Komura D, Ishikawa S. Machine learning approaches for pathologic diagnosis. Virchows Arch. 2019;475:131-8. https://doi.org/10.1007/s00428-019-02594-w
Rezende MT, Tobias AHG, Silva R, Oliveira P, Sombra de Medeiros F, Ushizima D, et al. CRIC Cervix Cell Classification. Fecha de consulta: 15 de junio de 2022. Disponible en: https://doi.org/10.6084/m9.figshare.c.4960286.v2
Nayar R, Wilbur DC, editors. The Bethesda System for reporting cervical cytology. 3rd edition. Chicago, IL: Springer; 2015. https://doi.org/10.1007/978-3-319-11074-5
Kräter M, Abuhattum S, Soteriou D, Jacobi A, Krüger T, Guck J, et al. AIDeveloper: Deep learning image classification in life science and beyond. Adv Sci. 2021;8:2003743. https://doi.org/10.1002/advs.202003743
Huang G, Liu Z, van Der Maaten L, Weinberger KQ. Densely connected convolutional networks. arXiv:1608.06993. 2017:4700-8. https://doi.org/10.48550/arXiv.1608.06993
Szegedy C, Liu W, Jia Y, Sermanet P, Reed S, Anguelov D, et al. Going deeper with convolutions. arXiv:1409.4842. 2015:1-9. https://doi.org/10.48550/arXiv.1409.4842
Howard AG, Zhu M, Chen B, Kalenichenko D, Wang W, Weyand T, et al. Mobilenets: Efficient convolutional neural networks for mobile vision applications. arXiv [Preprint]. 2017. arXiv:1704.04861.
Simonyan K, Zisserman A. Very deep convolutional networks for large-scale image recognition. arXiv [Preprint]. 2014. arXiv:1409.1556.
Hosna A, Merry E, Gyalmo J, Alom Z, Aung Z, Azim MA. Transfer learning: A friendly introduction. J Big Data. 2022;9:102. https://doi.org/10.1186/s40537-022-00652-w
Ministerio de Salud de Colombia. Guía de práctica clínica para la detección y manejo de lesiones precancerosas de cuello uterino. Bogotá, D. C.: Minsalud; 2014. p. 26-34.
Nanda K, McCrory DC, Myers ER, Bastian LA, Hasselblad V, Hickey JD, et al. Accuracy of the Papanicolaou test in screening for and follow-up of cervical cytologic abnormalities: A systematic review. Ann Intern Med. 2000;132:810-9. https://doi.org/10.7326/0003-4819-132-10-200005160-00009
Cao L, Yang J, Rong Z, Li L, Xia B, You C, et al. A novel attention-guided convolutional network for the detection of abnormal cervical cells in cervical cancer screening. Med Image Anal. 2021;73:102197. https://doi.org/10.1016/j.media.2021.102197
Sornapudi S, Brown GT, Xue Z, Long R, Allen L, Antani S. Comparing deep learning models for multi-cell classification in liquid-based cervical cytology images. AMIA Annu Symp Proc. 2019;2019:820-7. https://doi.org/10.48550/arXiv.1910.00722
Wang CW, Liou YA, Lin YJ, Chang CC, Chu PH, Lee YC, et al. Artificial intelligence-assisted fast screening cervical high grade squamous intraepithelial lesion and squamous cell carcinoma diagnosis and treatment planning. Sci Rep. 2021;11:16244. https://doi.org/10.1038/s41598-021-95545-y
Tang H, Cai D, Kong Y, Ye H, Ma Z, Lv H, et al. Cervical cytology screening facilitated by an artificial intelligence microscope: A preliminary study. Cancer Cytopathol. 2021;129:693-700. https://doi.org/10.1002/cncy.22425
Zhu XH, Li XM, Zhang WL, Liao MM, Li Y, Wang FF, et al. Clinical application of deep learning based artificial intelligence-assisted system in cervical cytology screening. Zhonghua Bing Li Xue Za Zhi. 2021;50:333-8. https://doi.org/10.3760/cma.j.cn112151-20201013-00780
Some similar items:
- Guillermo Terán-Ángel, Vestalia Rodríguez, Rosilved Silva, Olga Zerpa, Henk Schallig, Marian Ulrich, Maira Cabrera, Herramientas no invasivas en Venezuela: comparación entre las pruebas inmunoserológicas DAT, rK26 y rK39 en el diagnóstico de leishmaniasis visceral , Biomedica: Vol. 30 No. 1 (2010)
- Oscar F. Herrán, María F. Ardila, Martha P. Rojas, Gustavo A. Hernández, Design of dietary questionnaires to study the relationships between diet and cancer prevalence in Colombia , Biomedica: Vol. 30 No. 1 (2010)
- Libia M. Rodríguez, Mabel C. Giraldo, Natalia García, Laura Velásquez, Sara C. París, Cristiam M. Álvarez, Luis F. García, Human leucocyte antigen gene (HLA-A, HLA-B, HLA-DRB1) frequencies in deceased organ donors , Biomedica: Vol. 27 No. 4 (2007)
- Guillermo Terán-Angel, Henk Schallig, Olga Zerpa, Vestalia Rodríguez, Marian Ulrich, Maira Cabrera, The direct agglutination test as an alternative method for the diagnosis of canine and human visceral leishmaniasis , Biomedica: Vol. 27 No. 3 (2007)
- Juan Carlos Cataño, Eaton-Lambert myasthenic syndrome , Biomedica: Vol. 30 No. 3 (2010)
- Juan Carlos Herrera, Luis Fernando Isaza, José Luis Ramírez, Gonzalo Vásquez, Carlos Mario Muñetón, Detection of chromosome 17 aneuplody and TP53 gene deletion in a broad variety of solid tumors by dual-color fluorescence in situ hybridization (FISH) , Biomedica: Vol. 30 No. 3 (2010)
- Yenny M. Montenegro-Medina, Luz Aída Rey-Caro, Jurg Niederbacher, Ruth Aralí Martínez-Vega, Fredi Alexander Díaz-Quijano, Luis Ángel Villar-Centeno, Roll of antibodies antiplatelets in viral infection: a systematic review of literature , Biomedica: Vol. 31 No. 1 (2011)
- Yaliana Tafurt-Cardona, Leidy D. Jaramillo-Ruiz, Wilson Muñoz-Ordóñez, Sulma L. Muñoz-Benítez, Carlos H. Sierra-Torres, High frequency of chromosome aberrations observed in lymphocytes in postmenopausal obese women , Biomedica: Vol. 32 No. 3 (2012)
- Edgar Martínez-Salazar, Alberto Tobón-Castaño, Silvia Blair, Natural Plasmodium knowlesi malaria infections in humans , Biomedica: Vol. 32 (2012): Suplemento 1, Malaria
- Ricardo Cendales, Constanza Pardo, Claudia Uribe, Guillermo López, María Clara Yépez, Luis Eduardo Bravo, Data quality at population-based cancer registries in Colombia , Biomedica: Vol. 32 No. 4 (2012)
Copyright (c) 2025 Biomedica

This work is licensed under a Creative Commons Attribution 4.0 International License.
| Article metrics | |
|---|---|
| Abstract views | |
| Galley vies | |
| PDF Views | |
| HTML views | |
| Other views | |