Representaciones basadas en aprendizaje profundo como apoyo del diagnóstico de la COVID-19 en cortes de tomografía computarizada
Resumen
Introducción. La enfermedad por coronavirus (COVID-19) es actualmente el principal problema de salud pública en el mundo. En este contexto, el análisis automático de tomografías computarizadas (TC) surge como una herramienta diagnóstica complementaria que permite caracterizar hallazgos radiológicos, y categorizar y hacer el seguimiento de pacientes con COVID-19. Sin embargo, este análisis depende de la experiencia de los radiólogos, por lo que las valoraciones pueden ser subjetivas.
Objetivo. Explorar representaciones de aprendizaje profundo entrenadas con cortes de TC torácica para diferenciar automáticamente entre los casos de COVID-19 y personas no infectadas.
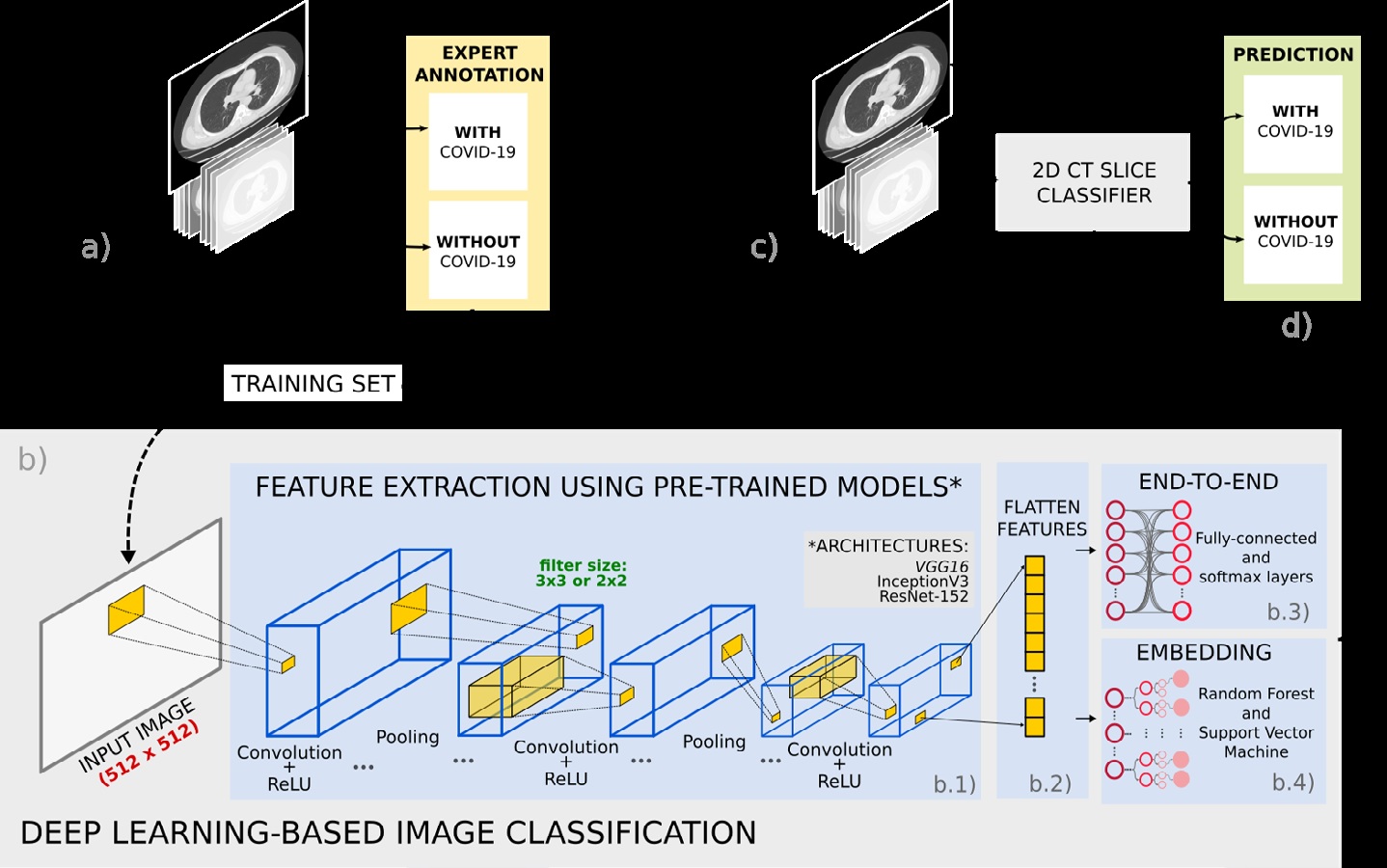
Materiales y métodos. Se usaron dos conjuntos de datos de TC: de SARS-CoV-2 CT (conjunto 1) y de la clínica FOSCAL (conjunto 2). Los modelos de aprendizaje supervisados y previamente entrenados en imágenes naturales, se ajustaron usando aprendizaje por transferencia. La clasificación se llevó a cabo mediante aprendizaje de extremo a extremo y clasificadores tales como los árboles de decisiones y las máquinas de soporte vectorial, alimentados por la representación profunda previamente aprendida.
Resultados. El enfoque de extremo a extremo alcanzó una exactitud promedio de 92,33 % (89,70 % de precisión) para el conjunto 1 y de 96,99 % (96,62 % de precisión) para el conjunto-2. La máquina de soporte vectorial alcanzó una exactitud promedio de 91,40 % (precisión del 95,77 %) para el conjunto-1 y del 96,00 % (precisión del 94,74 %) para el conjunto 2.
Conclusión. Las representaciones profundas lograron resultados sobresalientes al caracterizar patrones radiológicos usados en la detección de casos de COVID-19 a partir de estudios de TC y demostraron ser una potencial herramienta de apoyo del diagnóstico.
Descargas
Referencias bibliográficas
Guarner J. Three emerging coronaviruses in two decades: The story of SARS, MERS, and now COVID-19. Am J Clin Pathol. 2020;153:420-1. https://doi.org/10.1093/ajcp/aqaa029
Liu R, Han H, Liu F, Lv Z, Wu K, Liu Y, et al. Positive rate of RT-PCR detection of SARSCoV-2 infection in 4880 cases from one hospital in Wuhan, China, from Jan to Feb 2020. Clin Chim Acta. 2020;505:172-5. https://doi.org/10.1016/j.cca.2020.03.009
Johns Hopkins University & Medicine. COVID-19 Dashboard by the Center for Systems Science and Engineering (CSSE) at Johns Hopkins University (JHU). Accessed: June 5, 2021. Disponible en: https://coronavirus.jhu.edu/map.html
Wiersinga WJ, Rhodes A, Cheng AC, Peacock SJ, Prescott HC. Pathophysiology, transmission, diagnosis, and treatment of coronavirus disease 2019 (COVID-19): a review. JAMA. 2020;324:782-93. https://doi.org/10.1001/jama.2020.12839
Li L, Qin L, Xu Z, Yin Y, Wang X, Kong B, et al. Artificial intelligence distinguishes COVID-19 from community acquired pneumonia on chest CT. Radiology. 2020. https://doi.org/10.1148/radiol.2020200905
Aleta A, Martín-Corral D, Piontti AP, Ajelli M, Litvinova M, Chinazzi M, et al. Modelling the impact of testing, contact tracing and household quarantine on second waves of COVID-19. Nat Hum Behav. 2020;4:964-71. https://doi.org/10.1038/s41562-020-0931-9
Wang W, Xu Y, Gao R, Lu R, Han K, Wu G, et al. Detection of SARS-CoV-2 in different types of clinical specimens. JAMA. 2020;323:1843-4. https://doi.org/10.1001/jama.2020.3786
Kucirka LM, Lauer SA, Laeyendecker O, Boon D, Lessler J. Variation in false-negative rate of reverse transcriptase polymerase chain reaction–based SARS-CoV-2 tests by time since exposure. Ann Intern Med. 2020;173:262-7. https://doi.org/10.7326/M20-1495
Y. Kortela E, Kirjavainen V, Ahava MJ, Jokiranta ST, But A, Lindahl A, et al. Real-life clinical sensitivity of SARS-CoV-2 RT-PCR test in symptomatic patients. PLoS ONE. 2021;16:e0251661. https://doi.org/10.1371/journal.pone.0251661
Liang LL, Tseng CH, Ho HJ, Wu CY. Covid-19 mortality is negatively associated with test number and government effectiveness. Sci Rep. 2020;10:12567. https://doi.org/10.1038/s41598-020-68862-x
Inui S, Fujikawa A, Jitsu M, Kunishima N, Watanabe S, Suzuki Y, et al. Chest CT findings in cases from the cruise ship “Diamond Princess” with coronavirus disease 2019 (COVID-19). Radiol Cardiothorac Imaging. 2020;2:e200110. https://doi.org/10.1148/ryct.2020200110
Dai Wc, Zhang Hw, Yu J, Xu Hj, Chen H, Luo Sp, et al. CT imaging and differential diagnosis of COVID-19. Can Assoc Radiol J. 2020;71:195-200. https://doi.org/ 10.1177/0846537120913033
Fang Y, Zhang H, Xie J, Lin M, Ying L, Pang P, et al. Sensitivity of chest CT for COVID-19: comparison to RT-PCR. Radiology. 2020;296:E115-7. https://doi.org/10.1148/radiol.2020200432
Hope MD, Raptis CA, Shah A, Hammer MM, Henry TS. A role for CT in COVID-19? What data really tell us so far. Lancet. 2020;395:1189-90. https://doi.org/10.1016/S0140-6736(20)30728-5
Bai HX, Hsieh B, Xiong Z, Halsey K, Choi JW, Tran TML, et al. Performance of radiologists in differentiating COVID-19 from viral pneumonia on chest CT. Radiology. 2020;296:E46-54 https://doi.org/10.1148/radiol.2020200823
Xiao AT, Tong YX, Zhang S. False-negative of RT-PCR and prolonged nucleic acid conversion in COVID-19: Rather than recurrence. J Med Virol. 2020;92:1755-6. https://doi.org/10.1002/jmv.25855
Mahomed N, van Ginneken B, Philipsen RH, Meléndez J, Moore DP, Moodley H, et al. Computer-aided diagnosis for World Health Organization-defined chest radiograph primaryendpoint pneumonia in children. Pediatr Radiol. 2020;50:482-91. https://doi.org/10.1007/s00247-019-04593-0
Kermany DS, Goldbaum M, Cai W, Valentim CC, Liang H, Baxter SL, et al. Identifying medical diagnoses and treatable diseases by image-based deep learning. Cell. 2018;172:1122-31. https://doi.org/10.1016/j.cell.2018.02.010
Silva P, Luz E, Silva G, Moreira G, Silva R, Lucio D, et al. COVID-19 detection in CT images with deep learning: A voting-based scheme and cross-datasets analysis. Inform Med Unlocked. 2020;20:100427. https://doi.org/10.1016/j.imu.2020.100427
Ragab DA, Attallah O. FUSI-CAD: Coronavirus (COVID-19) diagnosis based on the fusion of CNNs and handcrafted features. PeerJ Comput Sci. 2020;6:e306. https://doi.org/10.7717/peerj-cs.306
Deng J, Dong W, Socher R, Li LJ, Li K, Fei-Fei L. ImageNet: A large-scale hierarchical image database. IEEE Conference on Computer Vision and Pattern Recognition. 2009. https://doi.org/10.1109/CVPR.2009.5206848
Soares E, Angelov P, Biaso S, Froes MH, Abe DK. SARS-CoV-2 CT-scan dataset: A large dataset of real patients CT scans for SARS-CoV-2 identification. medRxiv. 2020. https://doi.org/10.1101/2020.04.24.20078584
Hansell DM, Bankier AA, MacMahon H, McLoud TC, Muller NL, Remy J. Fleischner Society: Glossary of terms for thoracic imaging. Radiology. 2008;246:697-722. https://doi.org/10.1148/radiol.2462070712
Pan F, Ye T, Sun P, Gui S, Liang B, Li L, et al. Time course of lung changes on chest CT during recovery from 2019 novel coronavirus (COVID-19) pneumonia. Radiology. 2020;295:715-21. https://doi.org/10.1148/radiol.2020200370
Bernheim A, Mei X, Huang M, Yang Y, Fayad ZA, Zhang N, et al. Chest CT findings in coronavirus disease-19 (COVID-19): Relationship to duration of infection. Radiology. 2020;295:200463. https://doi.org/10.1148/radiol.2020200463
Parekh M, Donuru A, Balasubramanya R, Kapur S. Review of the chest CT differential diagnosis of ground-glass opacities in the COVID era. Radiology. 2020:297:E289-302. https://doi.org/10.1148/radiol.2020202504
Simonyan K, Zisserman A. Very deep convolutional networks for large-scale image recognition. arXiv preprint arXiv:14091556. The 3rd International Conference on Learning Representations, 2015. Disponible en: https://arxiv.org/abs/1409.1556
He K, Zhang X, Ren S, Sun J. Deep residual learning for image recognition. IEEE Conference on Computer Vision and Pattern Recognition. 2016. https://doi.org/10.1109/CVPR.2016.90
Szegedy C, Vanhoucke V, Ioffe S, Shlens J, Wojna Z. Rethinking the inception architecture for computer vision. IEEE Conference on Computer Vision and Pattern Recognition. 2016. https://doi.org/10.1109/CVPR.2016.308
Tan C, Sun F, Kong T, Zhang W, Yang C, Liu C. A survey on deep transfer learning. In: Kůrková V, Manolopoulos Y, Hammer B, Iliadis L, Maglogiannis I, editors. Artificial Neural Networks and Machine Learning – ICANN 2018. Lecture Notes in Computer Science, vol. 11141. Springer, Cham, Switzerland; 2018. https://doi.org/10.1007/978-3-030-01424-7_27
Hijazi S, Kumar R, Rowen C. Using convolutional neural networks for image recognition. San José, CA, USA: Cadence Design Systems Inc; 2015. p. 1-12.
Akilan T, Wu QJ, Jiang W. A feature embedding strategy for high-level CNN representations from multiple convnets. IEEE Global Conference on Signal and Information Processing. 2017. https://doi.org/10.1109/GlobalSIP.2017.8309150
Pham TD. A comprehensive study on classification of COVID-19 on computed tomography with pretrained convolutional neural networks. Sci Rep. 2020;10:1-8. https://doi.org/10.1038/s41598-020-74164-z
Alebiosu DO, Muhammad FP. Medical Image Classification: A Comparison of Deep Pretrained Neural Networks. IEEE Student Conference on Research and Development. 2019. https://doi.org/10.1109/SCORED.2019.8896277
Breiman L. Random forests. Machine Learning. 2001;45:5-32. https://doi.org/10.1023/A:1010933404324
Savas C, Dovis F. The impact of different kernel functions on the performance of scintillation detection based on support vector machines. Sensors. 2019;19:5219. https://doi.org/10.3390/s19235219
Amami R, Ayed DB, Ellouze N. Practical selection of SVM supervised parameters with different feature representations for vowel recognition. International Journal of Digital Content Technology and its Applications. 2013;7(9). https://doi.org/10.48550/arXiv.1507.06020
Algunos artículos similares:
- José Moreno-Montoya, El desafío de comunicar y controlar la epidemia por coronavirus , Biomédica: Vol. 40 Núm. 1 (2020)
- Juan Pimentel, Neil Andersson, Cloroquina y sus derivados en el manejo de la COVID-19: una revisión sistemática exploratoria , Biomédica: Vol. 40 Núm. Supl. 2 (2020): SARS-CoV-2 y COVID-19
| Estadísticas de artículo | |
|---|---|
| Vistas de resúmenes | |
| Vistas de PDF | |
| Descargas de PDF | |
| Vistas de HTML | |
| Otras vistas | |